In this section we will post cartoons representing relevant current and past key articles, manuscripts that are controversial or simply fun science. You will have one week to guess which publication the cartoon is representing. After that the title will be revealed including a short summary of the paper, points of relevance or consideration and important links for further reading
01/28/2019 Guess of the day
„ What do Banksy’s artwork & engram cells have in common?“ #MondayMotivation (hint: past week’s news is this week’s art)

Solution: Research published by Pignatelli, Ryan and colleagues in NeuroCellPress demonstrates how memory recall induces a transient increase in engram excitability. The cells can encode and switch between different memory functions, but how exactly is unknown. So, for a short period of time (~1h) we have an enhanced view on our surrounding, enhanced context recognition, potentially crucial for our survival. However, a prolonged excitability of the neurons is most likely unhealthy and this state thus passes.
To read more about this exciting research, see also the preview, actual paper or the Trinity College Dublin press release, which published some of our art along with the article.
3/13/2018 Guess the publication (6th ed.)

Solution: For this weeks’ Guess The Publication we did not only present a single cartoon, but a whole sci-art poster representing the concept, psychological mechanisms and neural basis of emotion regulation as presented in the synthetic review article by Kevin Ochsner, Jennifer Silvers and Jason Buhle (2012). Important to note is also, that this model was first proposed in 2002, elaborated on in 2005, further described as depicted here in 2012 and recently extended in a 2017 publication by Laura Braunstein, James Gross and Kevin Ochsner. The latest key features include a further distinction separating the nature of the emotion regulation goal (from implicit to explicit) and the nature of the emotion change process (from automatic to control). Such ongoing adaptations to the model of the cognitive control of emotions are made to reflect the increasing and changing knowledge gained through animal and human research studies.
But what is emotion regulation exactly? Emotion regulation describes the process of changing or controlling an emotion. This can either be an up- or down-regulation of the magnitude or duration of a feeling experienced. In order to understand emotion regulatory processes, it is helpful to first revisit steps included in the generation of an emotional response. In short, these include: a stimulus that is present in a certain context, attention directed at this stimulus, assigning a meaning to the stimulus and translating the stimulus into a bodily or mental representation. And so an emotional response is generated. However, not in all cases is it appropriate to immediately react on the first initial feeling generated. As for example, we may not have considered all the information available. And sometimes the feelings are warranted, but it is nevertheless socially inappropriate to immediately act on them. How well we are able to regulate our own emotions has been linked to personal well-being and the well-being of our surroundings. An inability to apply appropriate emotion regulation strategies has further been linked to psychiatric conditions, including conduct disorder, anxiety disorders or depression.
From a neuroimaging perspective, this review suggests that there is a network of brain areas activated by an emotional stimulus (including the amygdala, insula and ventral striatum), a network of brain regions performing the emotion regulation per se (including dorsomedial-, dorsolateral and ventrolateral prefrontal cortex, dorso-anterior cingulate cortex, posterior PFC and inferior parietal brain regions) and additional brain regions with a more intermediary role (including prefrontal and orbitofrontal cortex, superior/middle temporal gyrus and the temporoparietal junction). Additionally, as mediating pathways the dorsomedial or ventrolateral prefrontal regions are suggested to modulate amygdala reactivity through the ventromedial prefrontal cortex.
The here reviewed research and the proposed model of the cognitive control of emotions may lay the foundation for studies assessing all dimensions of emotion regulation behaviours in healthy or clinical populations. Disorder-specific patterns of emotion regulation abnormalities as observed in behavioural observations or neuronal assessments may hold the promise to inform about the relevant pathophysiology of mental disorders. Furthermore, they may provide indicators on treatment choice and/or explain treatment success. A current exemplary large-scale study focusing on emotion processing and regulation in psychiatric childhood disorders is FemNAT-CD, a multicentre neuroimaging study aiming to assess the genetic, behavioural and neurobiological markers of female conduct disorder.
Summary & Take Home Message: The here “cartoonized” review paper synthesizes functional imaging research on emotion regulation and proposes a basic model of the psychological mechanisms and neural systems involved. This evolving model of the cognitive control of emotions at all levels may lay the foundation for studies targeting the investigation not only in healthy, but also clinical populations.
Original Publication: Ochsner, K. N., Silvers, J. A., & Buhle, J. T. (2012). Functional imaging studies of emotion regulation: a synthetic review and evolving model of the cognitive control of emotion. Annals of the New York Academy of Sciences, 1251(1).
Fun Fact: At bornascientist.com, we do not only like Zombies – we are also Star Wars fans. For a lay language and fun description of emotion processing, regulation and the brain, read our publication in Frontiers for Young Minds called “Emotions and the brain or how to master the force”.
01/13/2018 Guess the publication (5th ed.)

Solution: Our latest cartoon of the week was inspired by the 2017 publication of Qiu and colleagues, published in Nature Human Behavior. This study questions a major challenge within the mainstream media or digital world to date: how can low quality information become widely popular and why do fake news eventually surpass the actual truth? Qiu and colleagues investigate whether the quality of a piece of information (i.e. how valid a statement is) is making it more or less likely to prevail.
Through their model, the authors weigh the relationship between the quality of and idea and the likelihood for it to go viral. While we may all wish to think that ultimately the truth would conquer over all other information available, this is unfortunately not the case. Two main reasons are identified which may contribute to the failing of the system’s ability to discriminate truth from random ideas or potentially dangerous fake news:
(1) Information overload (there is too much we have to process)
(2) Limited individual attention (we can only attend a limited amount of infos at once)
Social media allow their users to share large amount of “ideas” on a daily basis. But humans can only maintain a certain amount of social interactions during a single day. One consequence of this is that groups of people end up surrounded by a small circle, their own “bubble”, of friends. Such friends tend to share similar beliefs and mindsets, which consequently leads to a skewed flow of information in each person’s social media channel accelerating the problem even more.
Do you know how scientific results are obtained? Why can’t we simply conquer the mass of fake news online by providing equally as much science news? Research is the search for knowledge through the systematic and careful study of a topic, an object, or any source of information in order to test a hypothesis and gain new insight. We believe that once validated, the observation is proven to be correct with a high enough likelihood that it can be agreed on as being true by all. At least until proven differently. But by this definition science news can never be created so fast or in such volume as random ideas. In “Limited individual attention and online virality of low-quality information” Qiu and colleagues suggest that one way to overcome this problem is to control the use of bots that flood social media with low quality information.
Take Home Message: The quality of a piece of information is only weakly linked to its popularity. Qiu and colleagues highlight in their article how information overload and limited attention can lead to a system where quality of a piece of information or of an idea is not determining whether certain information do become popular on social networks or not. Digital misinformation has been ranked a major global risk for our society and it is important to question the high volume of misinformation observed online.
Original Publication: Qiu, X., Oliveira, D. F., Shirazi, A. S., Flammini, A., & Menczer, F. (2017). Limited individual attention and online virality of low-quality information. Nature Human Behaviour, 1(7), 0132.
Fun Fact: By the way, this is how @bornascientist would describe the path for reaching a scientific fact – through systematic and careful study and rigorous peer review. And in comparison the creation of a random idea or potential fake news:

11/24/2017 Guess the publication (4th ed.)

Solution: Our cartoon of the week was motivated by the 2017 publication of Gordon & Laumann and colleagues in Neuron, which was entitled “Precision functional mapping of individual human brains”. Nights filled with neuroimaging sessions organized by the group led to an unprecedented amount of hours’ worth of structural, functional and resting state MRI data in 10 adults. To be precise, 850 minutes of it for each individual. Not only did the team acquire all that data, they also provided it as a resource for others to further our understanding of individual human brain maps.
Let’s be honest, when looking at such individual brain maps (e.g. Fig 4. Graph Analysis within Gordon et al., 2017), they look stunning! And complicated. So, what exactly do they tell us again? The research team describes that one of the core ideas behind the start of their project was that there might be a limitation deriving from the fact that neuroimaging has focused predominantly on group-averaged brain data. This may have the disadvantage of losing detailed and more specific information from one individual and thus consequently reduces its clinical applicability. If neurobiological markers were to be used for prognostic purposes, they would have to be reliable (which refers to the consistency of a finding or how often the same result is obtained when re-evaluated) and valid (meaning that it has to correspond to real-world applications). The presented findings are a first indication for differences in individually obtained versus group-averaged network findings.
The midnight scan club and their work on individualized brain data is following pioneering work by scientists including Prof. Russell Poldrack from Stanford University. Poldrack provided a first approach on individual brain mapping, by scanning himself twice for close to 18 months. By doing so, it became the most in-depth studied individual human brain ever (read more in the following article, or see this original publication called “Long-term neural and physiological phenotyping of a single human” which appeared in Nature Communications). We could not help, but wonder – how did it feel for that brain to be scanned so often: like being a celebrity or more like being in the movie groundhog day?

But joking aside, these examples sure make us interested in knowing more about where the field moves next. It certainly has inspired many to look at data differently. Let’s see how much of a paradigm shift in data collection and interpretation will follow!
Take home message: An increased focus on more quality data from one individual may hold more specific and precise information as opposed to data obtained by a large group average. Instead of averaging data across a certain group, hours of scan data in each participant was obtained in order to increase specificity and precision within a single individual. Through a highly sampled and now publicly available ten-subject data set individual variations in the basic brain organization were revealed across all participants scanned (i.e. circular networks in some, linear networks in other individuals). Such approaches may hold promise to ultimately inform about brain networks in health and disease.
Fun fact: The term midnight scan club is reflective of the fact that the group used scan time after midnight in order to being able to scan for long periods of time and most importantly being able to pay for it (annotation: most scan sites charge >500CHF/USD per scan hour).
Original publication:
Gordon, Laumann, Gilmore, Newbold, Greene, Berg, Ortega, Hoyt-Drazen, Gratton, Sun, Hampton, Coalson, Nguyen, McDermott, Shimony, Snyder, Schlaggar, Petersen, Nelson, Dosenbach (2017). Precision Functional Mapping of Individual Human Brains. Neuron, 95 (4), 791-807.
11/02/2017 Guess the publication (3rd ed.)

Solution: This week’s Guess the Publication was inspired by the 2009 poster and later publication of Bennett and colleagues 1 . Their research team conducted an fMRI study using a dead Atlantic Salmon as the main research subject. No, we did not make this up. In fact, many everyday objects look quite beautiful when scanned using an MRI machine. But this story with the salmon was about more than admiring the amazing inner works of fish visualized by MRI machines. But what? Why would a group of skilled scientists invest time into scanning a dead fish? They would do so, because the real purpose of this experiment was to show that neural activation obtained by functional MRI can be spurious unless the appropriate correction methods are applied!
The problem of coincidental findings in neuroscience can result from a problem called multiple comparison problem. When analysing neuroimaging data using a whole-brain analysis approach we run a test for every voxel within this whole-brain mask. This means for every single one of tens of thousands of voxels a test is computed. As a result of running that many tests, the likelihood of getting a false positive finding increases drastically. The problem of multiple comparison can be visualized by the example of someone throwing a dice over and over again (see this youtube video for an easy description of the topic), until you receive the number you hope for. The more times you throw the dice, the more likely you get the number right. Or, in fMRI analysis, with every statistical test run in addition to the previous one (and we said it’s tens of thousands of tests), the likelihood of finding spurious activation increases. With every additional tested voxel in the brain. This means nothing else as that the likelihood of a certain result increases with the number of tries. There are different correction methods in neuroscience to overcome this problem. These methods for example include family-wise error correction (FWE) and the false detection rate (FDR). Both represent an expectancy, that part of the positive discoveries will be false positives. There are also new techniques available, including threshold free cluster enhancement techniques (TFCE), that might offer sensitivity while taking voxel- and cluster-based information into account. But how exactly each one of them work is another story.
To go back to our study – what does the “salmon-experiment” have to do with the topic of multiple comparisons? Bennett and colleagues showed, that at a threshold of p < 0.001 with no correction for multiple comparison it is possible to obtain neuronal brain activation, even in a dead salmon. These findings disappeared upon employing FWE or FDR corrections.
Take home message: Without proper correction methods you may be looking at nothing more than a red herring (or hot fishy air). Scientists spend so much time and effort on acquiring quality data in order to venture further than anyone before. How we handle that data afterwards is a big responsibility, especially, because of the chance of misinterpretation. And also, it is o.k. to think outside of the box – it could go a long way.
Fun fact: this study did win an Ig Nobel award. The Ig Nobel award is dedicated to research studies that first make you laugh, then think. The awards honours those thinking outside the box and we can only recommend to read through the awards page and recipient’s work. You can also read more about the dead salmon study through an entertaining blog post that appeared in Scientific American. If you do, you will understand that it is not a coincidence that there was a pumpkin in the trash bin of our drawing representing this study.
Original publication:
[1] Bennett, C. M., Baird, A. A., Miller, M. B., & Wolford, G. L. (2011). Neural correlates of interspecies perspective taking in the post-mortem atlantic salmon: an argument for proper multiple comparisons correction. Journal of Serendipitous and Unexpected Results, 1, 1-5.
10/09/2017 Guess the publication (2nd ed.)
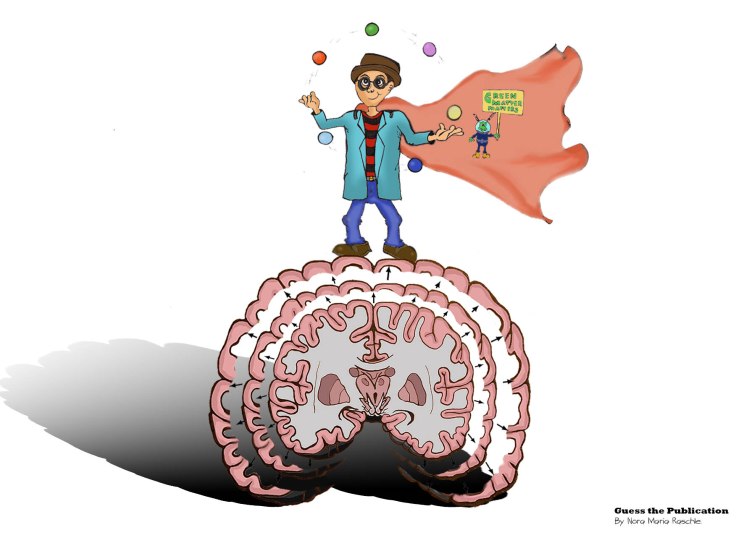
Solution: In the early 2000s, different studies using magnetic resonance imaging were able to show that the brain is adapting according to our behavior or experience 1,2,3. This week’s Guess the Publication is based on the 2004 study conducted by Draganski and colleagues3 entitled „Neuroplasticity: Changes in grey matter induced by training“. In their study the authors demonstrate stimulus-dependent change in the brain’s macroscopic structure, or anatomy, in participants that were learning a completely new skill, namely juggling. One important conclusion of the authors’ results lies in demonstrating, that no matter what age our brains still have the capability to change. More specifically, what we experience (environment and training) impacts brain anatomy. Brain development is thus in some ways a constant process and not something that is finalized at the end of late adolescence. Importantly, within their study design Draganski and his colleagues used a longitudinal approach that included healthy non-juggling adults that received three months of training (experimental/juggling group) and healthy non-juggling adults that received no training (control group). They scanned both a control and a juggling group before study start, after three months of practice for the jugglers, but not the controls, and then again three months later. In the last three months the juggler group also had to refrain from practicing (no training).

Before the juggling phase there were no statistically significant differences in the grey matter volume of the two groups. After just three months of practice the juggler group showed increased grey matter volume in motion-selective visual areas compared to the no-juggle control group. At the last scan (after three months with neither of the groups juggling), the expansion decreased in the juggler group, showing that the lack of practice was associated with a loss of the before acquired grey matter volume.
Take home message: No matter your age, if you are investing time in learning something new, this will impact your brain structure and function already after a few weeks. However, if you want to keep up those benefits, you have to continue practicing. Therefore, our main message of the week is: USE IT OR LOSE IT!
Original manuscript:
3 http://www.nature.com/nature/journal/v427/n6972/full/427311a.html?foxtrotcallback=true
Relevant reading:
1 http://www.pnas.org/content/97/8/4398.short
2 https://www.nature.com/articles/nrn843
09/28/2017 Guess the publication (1st ed.)
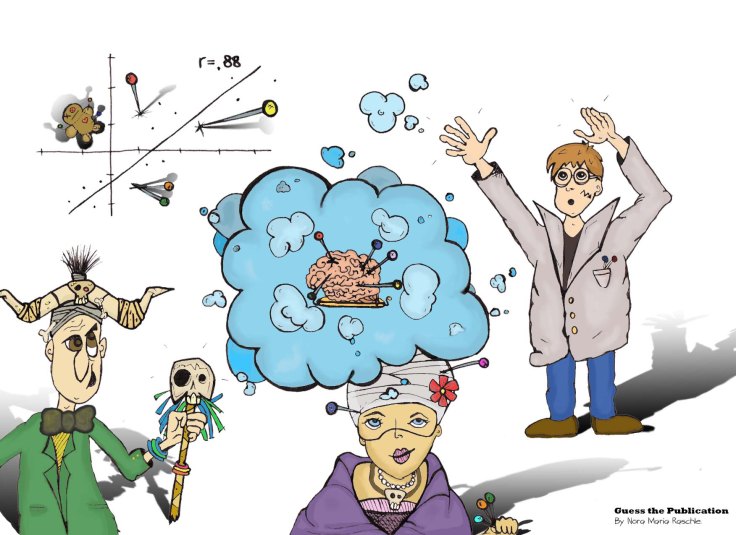
Solution: In their 2009 paper, originally called “Voodoo correlations in social neuroscience” and later renamed “Puzzlingly high correlations in fMRI studies of emotion, personality, and social cognition”, Vul and colleagues critically question the presence of impossibly high correlations in social neuroscience fMRI studies. While followed by many critical discussions about the correctness of assumptions and statistical arguments in their own paper (see for example 1, 2), one important contribution of this article to the field is bringing a focus on the problem that non-independence errors can lead to significant correlations out of noise. The most prominent form of a non-independence error is for example selection-bias. This happens when findings do result from correlations between voxels of certain brain areas and behaviour, when the behavioral variable was influencing the null hypothesis in the first places. However, despite the importance of the topic of non-independence errors, the manuscript was also considered problematic by some research teams due to (1) the use of the original title and general language in the manuscript (e.g. “voodoo correlations” immediately implies “fraudulence”) and (2) the inclusion of likewise wrong statistical arguments. Especially the strong negative connotations attributed to the whole field of social Neuroscience were reason to spark a lot of outcry post publication.
Take away message: When investigating a statistical relationship between a behavioural measure and voxels in the brain, you cannot apply a secondary non-independent test to that same set of voxels using the variable that you have already shown to distinguish the data. The resulting conclusion is biased.
Original manuscript: http://journals.sagepub.com/doi/abs/10.1111/j.1745-6924.2009.01125.x
Some critical reading related to this topic:
2 http://cogns.northwestern.edu/cbmg/replyVul.pdf
3 https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5017149/
4 http://www.jstor.org/stable/41613484?seq=1#page_scan_tab_contents
5 https://www.edvul.com/voodoocorr.php
6 http://neurocritic.blogspot.ch/2009/01/voodoo-correlations-in-social.html
7 http://www.ohbmbrainmappingblog.com/blog/keep-calm-and-scan-on
congrats! I tried same with money bank politics but my painter jumped off 🙂
see on twitter “above fog”
LikeLiked by 1 person
Hi Mike, thank you! And sorry to hear about your painter leaving the project. We do all illustrations and cartoons ourselves. Which might work in our favour here.
LikeLike